Automated Processing of Solid Tissues into Single Cells or Nuclei for Genomics and Cell Biology Applications with the Singulator™ 100 and 200 Systems
Download Our Poster Presented at PAG30
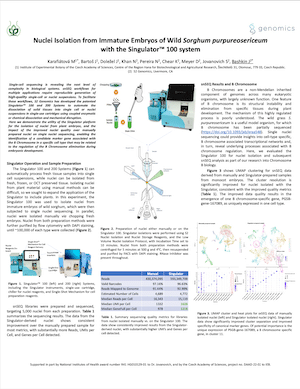
Single-cell sequencing workflows require reproducible generation of high-quality single-cell or -nuclei suspensions. Here, we demonstrate the utility of automated nuclei isolation from monocot plant embryos and the impact on the quality of single nuclei sequencing. The Singulator 100 platform was used to isolate nuclei from immature embryos of wild Sorghum purpureosericeum, which were then subjected to single nuclei sequencing. The improved data quality over manually prepared nuclei suspension enabled the identification of a candidate marker gene associated with the B chromosome in a specific cell type that may be associated with the regulation of the B chromosome elimination during embryonic development. The routine, automated isolation of high-quality nuclei from plant material can enable new avenues of research in genomics for botany.
This poster was presented at the Plant & Animal Genome Conference (PAG30).


